Grantor: Intelligence Advanced Research Projects Agency (IARPA)
Grant Program: Hybrid Forecasting Competition (HFC)
Start Date: 01 October 2018
End Date: 30 September 2021
Tufts PI: Elena Naumova
Forecasting is a systematic scaffolding of mathematical algorithms that use structural, dynamic, and iterative data to predict future events. Near-term, short-term, and long-term forecast models are essential in developing effective preventive strategies, resource allocation scenarios and early warning systems. The most effective forecasts are reliable (minimize prediction error), accurate (successful prediction of health outcome), complex (capturing outcome dynamics), valuable (actionable), clear (easily understood), sharp (robust across multiple conditions), and adaptive (accommodating real-time data).These models are ideally accommodate rapid changes in incoming data and contain sufficient complexity to capture disease etiology, seasonal behavior, and social dynamics. This project seeks to apply time series methods to improve the forecasting potential for infectious diseases or humanitarian emergencies.
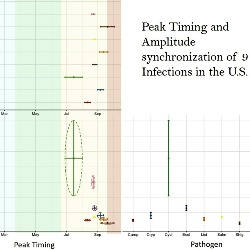
Foodborne Diseases
Modern food networks, from production to consumption, are vulnerable to foodborne infections that cause outbreaks difficult to track and control. Many foodborne infections are notorious for frequent summer outbreaks. We use a time series approach applied to FoodNet data and various visualization tools to detect periods of high and low foodborne disease incidence, identify high risk geographic areas and potential driving factors, explore synchronization of foodborne outbreaks, and design disease calendar to ease the communication of complex spatiotemporal patterns.
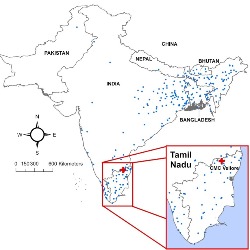
Cholera Outbreaks
Cholera is a preventable and treatable, yet potentially fatal bacterial disease that causes diarrhea and dehydration. We investigate cholera outbreaks in resource-scarce areas where disease surveillance is limited. We also focusing on regions where people are facing humanitarian emergencies due to natural disasters or war conflicts. We develop models to better understand factors effecting cholera transmission and intensity and design forecasts tailored to characteristics of places, times, and population at risk. We explore ways to better utilize routinely collected and maintained data streams, like electronic hospitalization records, laboratory testing reports, and environmental monitoring.
Influenza
The worldwide attention to influenza prevention has been resulted in increased availability of influenza surveillance data in the recent years. In 1997, WHO launched FluNet, the first web-based tool for global influenza virological surveillance. We are developing methods to examine factors that modify the timing and intensity of seasonal influenza in data-rich and data-limited settings. We are targeting countries with limited recourses or depleted economies due to natural disasters or humanitarian emergencies, where the quality and availability of reporting is still insufficient for developing reliable forecasts. Our research proposes a method to enable seasonal forecasts in data-sparse settings by triangulating temporal patterns observed in data-rich countries.
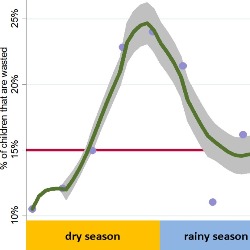
Famine
The USAID Famine Early Warning System (FEWS-NET) periodically assesses famine risk across several countries. However, FEWS-NET predictions are not updated in real-time and reports provide insufficient information on risk factors driving conditions of acute malnutrition. While famine has a technical definition, the declaration of famine often manifests a political process that may lack transparency. Our research investigates how a variety of socioeconomic risk factors can be used to develop near-term forecasts of food insecurity as a tool for early-warning famine declarations.