Current systems of interest are GI tract microbiota and industrial cell lines used for biologics production. To study these systems, we develop tools that can helps us systematically explore the chemical diversity of metabolites and quantify the activities of specific metabolic pathways.
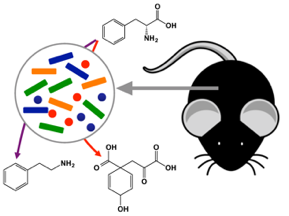
Microbiota metabolites – The human body harbors trillions of commensal microorganisms that interact with the host in a variety of ways to profoundly impact the host’s physiology in health and disease. A large concentration of these organisms is found in the GI tract. Well-known functions provided by the gut microbiota include digestion of complex nutrients and immune response to foodborne pathogens. A significant alteration of the microbiota, or dysbiosis, correlates with not only GI diseases, but also many other chronic diseases and developmental disorders that have an inflammatory component. Better understanding the mechanisms of host-microbiota interactions could unlock new ways to diagnose or treat these conditions. An important axis of communication between the host and gut microbiota utilizes bacterial metabolites that result from the catabolism of dietary residues or host-derived intestinal molecules such as mucin. Findings by researchers in our lab and other groups have shown that these metabolites activate specific host cellular receptors to modulate immune cell activity and inflammatory responses in the intestine, liver, and other tissues. These metabolites can deplete under dysbiosis, for example due to shift toward a Western diet or antibiotic exposure, disrupting the communication between host and microbiota. The goals of our research are to identify these bacterially derived metabolite ligands, elucidate the mechanisms of action, and determine how various environmental factors causing dysbiosis impact the metabolites’ source enzymes and organisms.
Clostridioides difficile is a pathogen that can infect the intestine. It is difficult to treat due to spore formation, leading to recurrence. We are investigating the role of bile acids, oligosaccharides, and other products of gut commensal bacteria in promoting or inhibiting the germination and outgrowth of C. difficile. We want to understand how broad spectrum antibiotics affect the population of bacteria that produce these metabolites and could potentially impact infection and recurrence.
Diet-derived natural products such as flavonoids have the potential to promote gut health by engaging host cellular receptors. We are interested in the consequences of metabolic activation (or deactivation) of these compounds by gut bacteria. We want to understand how the profile of bacterial enzymes in the intestine impacts the individual’s response to flavonoid enriched diets.
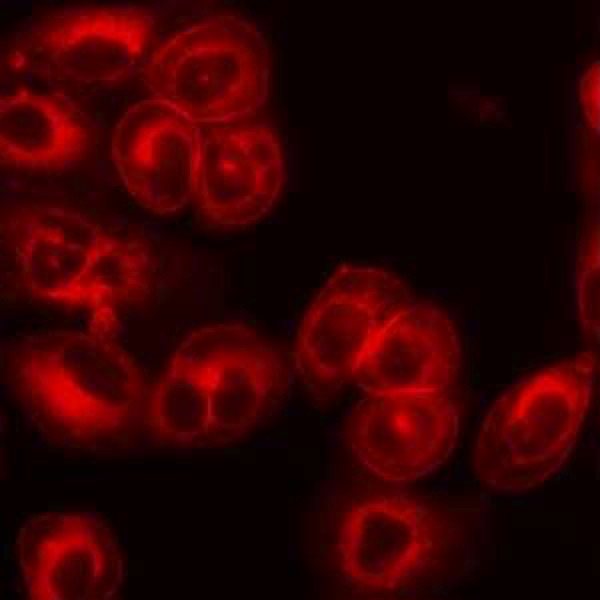
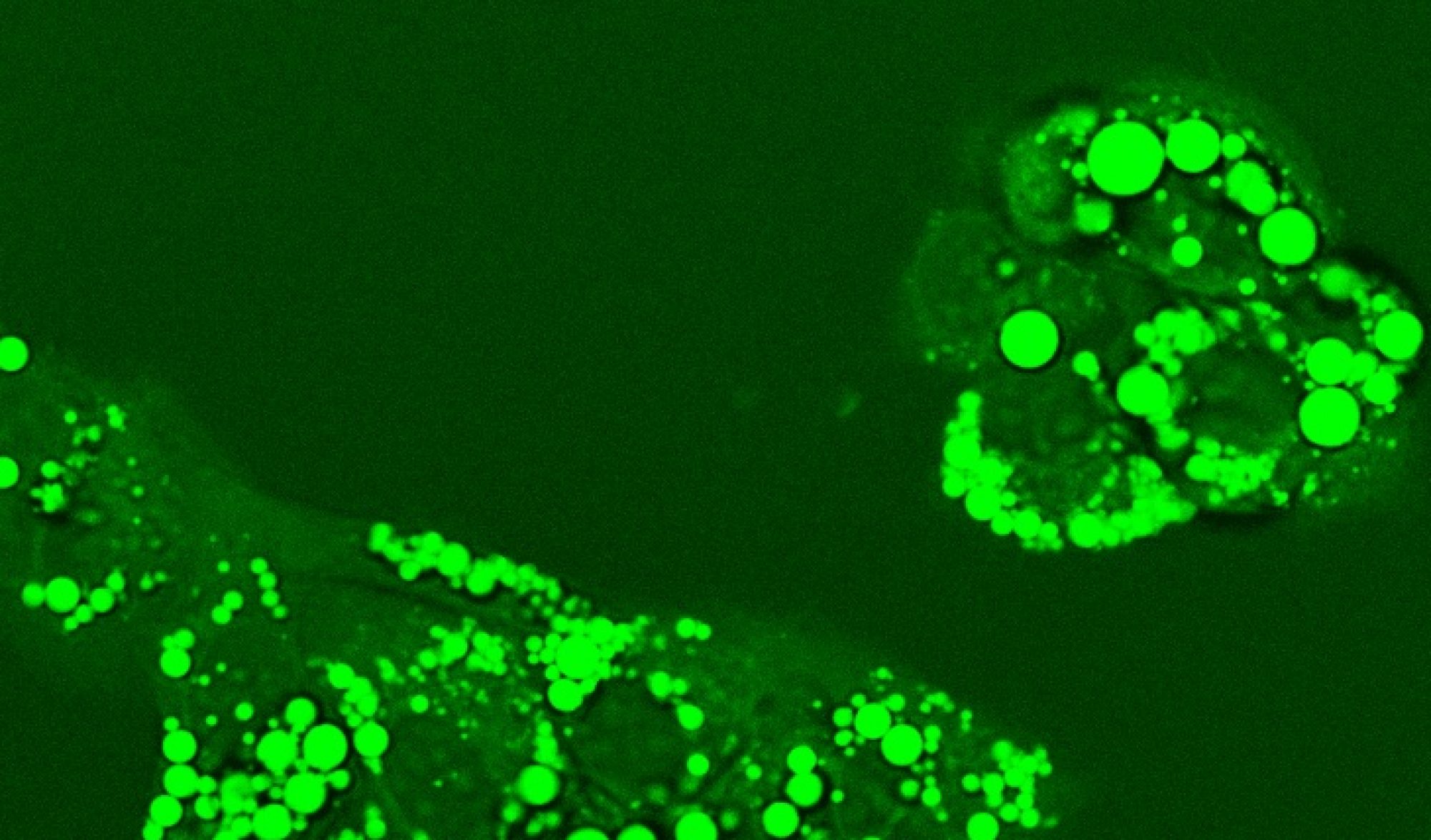
CHO cell metabolism – Chinese hamster ovary (CHO) cells are widely used for industrial production of biopharmaceuticals, especially monoclonal antibodies (mAbs), due to these cells’ capacity to support proper protein folding and post-translational modifications. Advances in process control, medium formulation, and host cell engineering have dramatically increased the volumetric productivity of CHO cell lines. As the CHO cells grow to very high cell densities in fed-batch systems, metabolic byproducts can accumulate to high concentrations. Well-known examples are lactate and ammonia, which can negatively impact culture performance. Recent research suggests that the number of significantly accumulating byproducts is large, but the byproducts remain largely uncharacterized. Moreover, the metabolic profile can vary dramatically across cell lines even when they are derived from the same host lineage. Better understanding how the byproducts are generated and what impact they have on cell physiology could lead to novel strategies for enhancing CHO cell-based bioprocess performance. In collaboration with industrial partners, we are characterizing CHO cell metabolic byproducts as potential indicators of metabolic inefficiency and investigating the implications of inefficiency on cell growth and protein biosynthesis.
Tools for metabolomics – Using targeted LC-MS assays, we can obtain quantitative measurements on metabolites to estimate the activity of a pathway or determine a physiologically relevant range of doses for functional characterization. The LC-MS experiments can also be performed in an untargeted fashion to broadly profile a significant fraction of all metabolites in biological system of interest. These untargeted experiments can detect thousands of compounds, which presents a challenge in analyzing the data. Typically, only a small subset of these compounds can be annotated with chemical identities, limiting data interpretation. We are working to improve the efficiency and accuracy of meabolomics data annotation by developing computational tools that model the biological context of samples and analyze the correlations between detected compounds.